Abbreviations: AF Alexa Fluor ATG autophagy related BafA1 bafilomycin A1 BSA bovine serum albumin DAPI 4,6-diamidino-2-phenylindole DMEM Dulbecco’s modified Eagle’s medium DMSO dimethyl sulfoxide EDTA ethylenediaminetetraacetic acid EBSS Earle’s balanced salt solution FBS fetal bovine serum GFP green fluorescent protein LD lipid droplet LSM laser scanning microscope MAP1LC3B microtubule associated protein 1 light chain 3 beta MTOR mechanistic target of rapamycin kinase PBS phosphate-buffered saline PIK3C3/VPS34 phosphatidylinositol 3-kinase catalytic subunit type 3 SQSTM1 sequestosome 1 TIFF tagged image file format U2OS U-2 OS cell line WIPI WD repeat domain, phosphoinositide interacting 
All protocols and software pipelines can be quickly and easily adapted for the use of alternative autophagy markers or cell types, and can also be used for high-throughput purposes. In addition, we provide CellProfiler pipelines for endogenous SQSTM1/p62 (sequestosome 1) or intracellular lipid droplet (LD) analysis, suitable to assess forms of selective autophagy. 
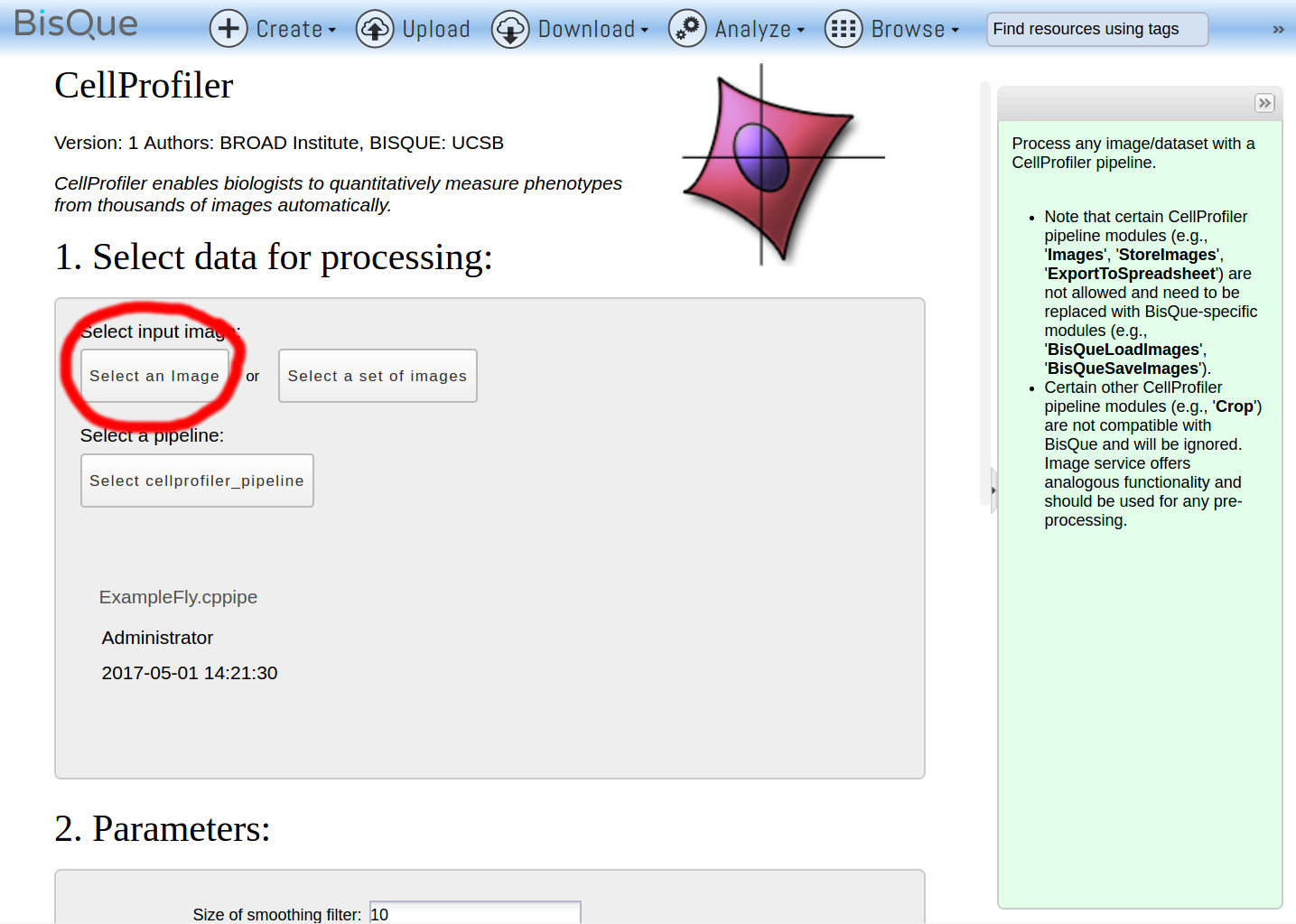
The CellProfiler protocol is provided as a ready-to-use software pipeline, and the creation of this pipeline is detailed in both text and video formats. Here we present a method for open source CellProfiler software-based analysis for quantitative autophagy assessments using GFP-tagged WIPI1 (WD repeat domain, phosphoinositide interacting 1) images acquired with Airyscan or confocal laser-scanning microscopy. In this context, an automated analysis of the number and size of recognized puncta is preferable to a manual count, because more reliable results can be generated in a short time. Basic Protocol 1: Studying cell morphology and cell migration in time-lapse datasets using TrackMate (Fiji) and CellProfiler Basic Protocol 2: Creating whole plate montages to easily assess adaptability of segmentation parameters.ĬellProfiler ImageJ cell tracking image stitching segmentation.Single cell-based analysis of macroautophagy/autophagy is largely achieved through the use of fluorescence microscopy to detect autophagy-related proteins that associate with autophagic membranes and therefore can be quantified as fluorescent puncta. ImageJ and CellProfiler are both committed to interoperability between their platforms, with ongoing development to improve how both are leveraged from the other. While both programs can be and are often used separately, these pipelines demonstrate the benefits of using them together for image analysis workflows. No single platform can provide all the key and most efficient functionality needed for all studies. Here, we share two pipelines demonstrating mechanisms for productively and conveniently integrating ImageJ and CellProfiler for (1) studying cell morphology and migration via tracking, and (2) advanced stitching techniques for handling large, tiled image sets to improve segmentation. Although many image analysis problems can be well solved with one or the other, using these two platforms together in a single workflow can be powerful. ImageJ's traditional strength is in single-image processing and investigation, while CellProfiler is designed for building large-scale, modular analysis pipelines. ImageJ and CellProfiler have long been leading open-source platforms in the field of bioimage analysis.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
